Research
News from the group:
Research Exchange Fellowships - IAESTE (apply)
CAMDA 2023 -
ISMB Conference Track, 26-27 July, Lyon, France (read more)
World-leading patient stratification - graph based cancer data integration (read more)
Confirming molecular mechanisms of tendon regeneration - a powerful ovine fetal model (read more)
Sequencing Quality Control
(SEQC) project,
MAQC Consortium 2011–2014 (read more)
MAQC Consortium 2011–2014 (read more)
Host–parasite interactions in biocontrol, WWTF grant 2010–2013
(read more)
Power and limitations of RNA-Seq,
FDA SEQC, Nature Biotechnology (read more)
FDA SEQC, Nature Biotechnology (read more)
Characterization and improvement of RNA-Seq precision,
Bioinformatics (read more)
Bioinformatics (read more)
Impact of heavy tails in microarray
analysis,
Bioinformatics
(read more)
Novel conserved repeats in sorting signals,
FEBS Journal (read more)
FEBS Journal (read more)
RNA interference in ageing
research,
Gerontology (read more)
Gerontology (read more)
Modelling Thermodymamics of Microarray Hybridization
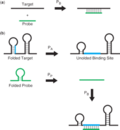 Ulrike Mï¿œckstein is
studying the application of RNA and DNA folding models for the
prediction of microarray probe hybridization behaviour.
Ulrike Mï¿œckstein is
studying the application of RNA and DNA folding models for the
prediction of microarray probe hybridization behaviour.
Partners
 We are grateful for the support of the structural modelling
experts Ivo Hofacker and his colleagues
at the Theoretical
Biochemistry Group of the University of Vienna.
We are grateful for the support of the structural modelling
experts Ivo Hofacker and his colleagues
at the Theoretical
Biochemistry Group of the University of Vienna.
Funding
 Support for this project was won through competition for the
Austrian Academy of Sciences Anniversary Fund of the City of
Vienna
Support for this project was won through competition for the
Austrian Academy of Sciences Anniversary Fund of the City of
Vienna


